What does under or over-dispersion look like?
One issue that often comes up in analysis with linear models is under or over dispersion.
For instance, if you are fitting a linear regression model, you are assuming the residuals (difference between the line of best fit and the data-points) are normally distributed. Residuals are said to be overdispersed if they are ‘fatter’ in the tails than a normal bell curve. Whereas, if the residuals are too peaked in the middle, they are said to be under-dispersed.
Under or over dispersion is an issue because it can bias the calculation of p-values. Over-dispersion is often of particular concern because it may cause p-values that are biased too low. If you are taking 0.05 as significant, over-dispersion will mean you would reject the null hypothesis more than 5% of the time. Thus, you may be more likely to falsely reject the null hypothesis.
If there is under-dispersion the opposite is true, your p-values may be too high. P-values that are biased high can be a problem, because they will give you low power to detect real effects.
So, let’s take a look at under and overdispersion. First, up here is a normal bell curve:
x <- seq(-4, 4, length.out = 100)
p <- dnorm(x, mean = 0, sd = 1)
plot(x, p, type = 'l', lwd = 2, col = "red",
xlab = "Residual", ylab = "Density")
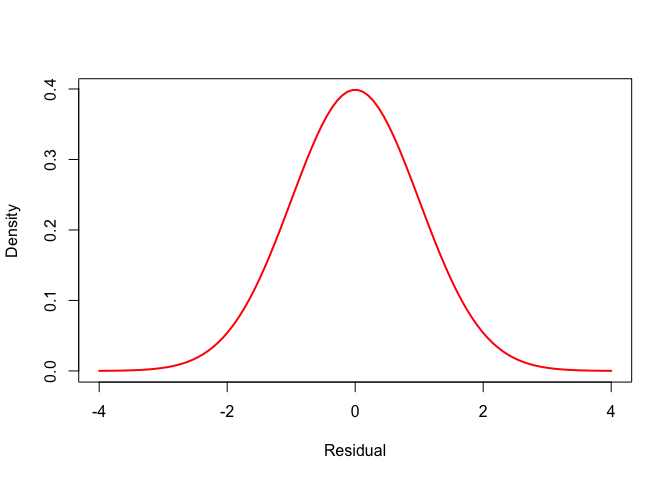
Now, we can add an over-dispersed curve to that. Here is one, calculated using the Student t distribution:
p_t <- dt(x, df = 1.4, ncp = 0)
plot(x, p, type = 'l', lwd = 2, col = "red",
xlab = "Residual", ylab = "Density")
lines(x, p_t, col = "darkblue", lwd = 2)
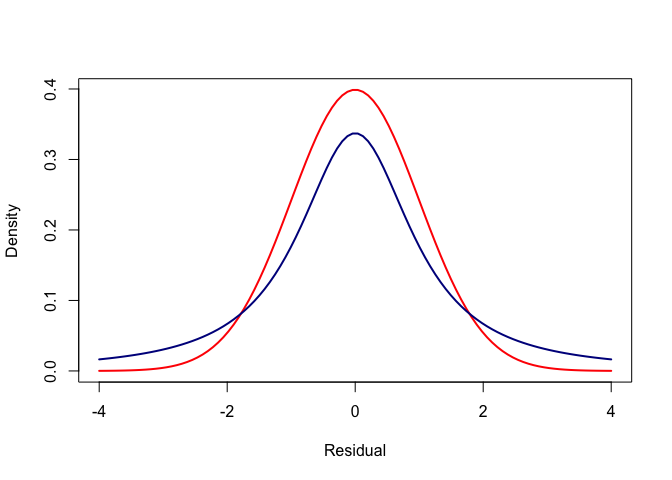
It wouldn’t matter how large we made the standard devation of the normal curve, we would never get it to match the Student-t.
Finally, let’s draw an under-dispersed distribution, using the Laplace distribution
library(rmutil)
p_l <- dlaplace(x, m = 0, s = 0.4)
Now plot them all together:
plot(x, p_l, xlim = c(-4, 4), lwd = 2, xlab = "Residuals", main = "", type = "l")
lines(x, p, col = "red", lwd = 2, lty = 2)
lines(x, p_t, col = "darkblue", lwd = 2)
legend("topright", legend = c("normal", "under-dispersed", "over-dispersed"),
lty = c(2,1,1), col = c("red", "black", "darkblue"), lwd = 2)
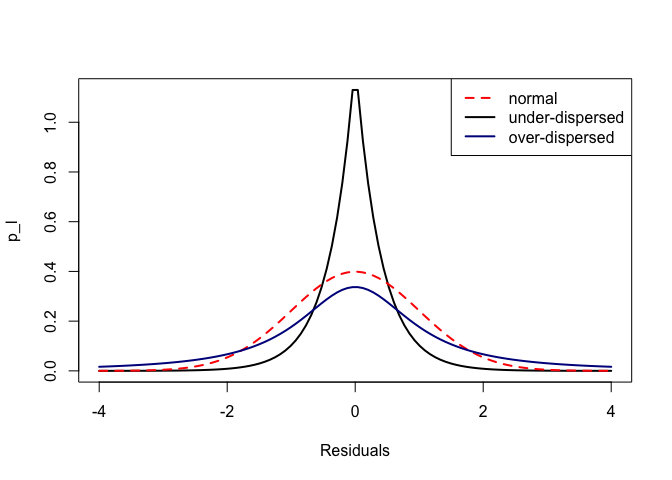
Here’s what they look like if we sample measures from each distribution and plot them on a normal QQ plot, which you may be familiar with. It is one of the standard checks of model residuals:
set.seed(1997)
par(mfrow = c(1,3))
qqnorm(rnorm(1000, mean = 0, sd = 1), main = "normal")
abline(0,1)
qqnorm(rt(1000, df = 1.4, ncp = 0), main = "over-dispersed")
abline(0,1)
qqnorm(rlaplace(1000, m = 0, s = 0.4), main = "under-dispersed")
abline(0,1)
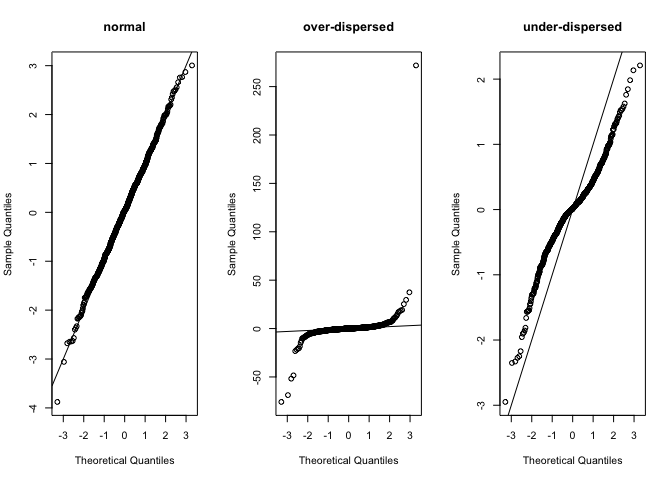
Playing around with the parameters for each distribution should help your understanding. For instnace, if you set the scale for the Laplace distribution to a larger number it will become over-dispersed, because it gets fatter tails than the normal (despite being more peaked at its mode).
Under-dispersion is more common than you might think. It can occur when you have a censoring process. For instance, perhaps you machine can only measure length’s to a certain precision and any distance that is to small gets rounded down to zero.